eXtensions
Macs in Science |

Benjamin Khoo began the day by outlining Apple's innovations before focusing on Science and the Mac. As 41% of Life Sciences (e.g. DNA; Proteomics) use Macs, there is already widespread use of the platform. It is particularly valuable as Mac clients, servers and clusters all use OS X.
He highlighted the MacBookPro, a strong contender for fieldwork, and the highly expandable MacPro (now with an 8-core processor): with its PCI Express slots it can support up to eight displays. A major advantage of the platform is that OS X for the server comes with unlimited licencing, unlike other operating systems.
Benjamin showed slides of the Virginia Tech. installation of Mac clusters which at one time was one of the world's fastest supercomputers. The software for clustering is built in to all Macs: it just has to be turned on. As an example, he told us about a small cluster he had constructed using four Mac minis.
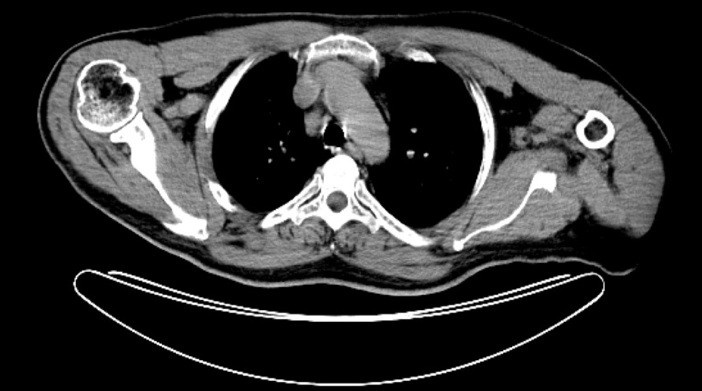
 Louis Lou, from Beijing, built on Benjamin's introduction in his focus on medical imaging, particularly the rise in 3D graphics use, like MRI and CT scans. During his peresentation, he looked at software available in this specialised field: Osirix, Resolution MD, Atamai, Volume Graphics, HeartIT, and NeuroLens. Osirix is available from the Apple website as a 35MB download.
Louis Lou, from Beijing, built on Benjamin's introduction in his focus on medical imaging, particularly the rise in 3D graphics use, like MRI and CT scans. During his peresentation, he looked at software available in this specialised field: Osirix, Resolution MD, Atamai, Volume Graphics, HeartIT, and NeuroLens. Osirix is available from the Apple website as a 35MB download.
Osirix, is Open Source software and therefore free. It was designed for radiologists to assist better workflows, from the scanner to the workstation, reports to the radiologist and from there to the physician or specialist. He cited examples of the way graphics-handling Macs were being integrated into some of China's hospitals.
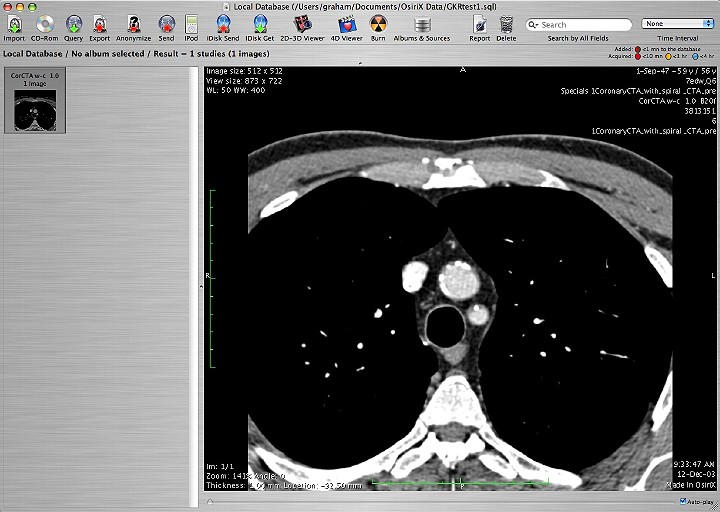
It was quite clear that this type of file handling was of interest to many attending. This was particularly evident when Louis displayed a fusion of image types that Osirix made possible.
One of the imaging applications that was of particular interest here was HeartIT, which allows high quality images to be displayed in a browser.
Louis enlarged on the Mac's flexibility and explained how data could be shared using iChat and its multi-person conferencing ability: useful for consultation or for distance teaching .
After lunch, Darran Nathan of Progeniq outlined a programmable processor that was valuable in accelerating computer-intensive applications particularly when large amounts of data with repetitive calculations were being used. An example might be the processsing required when DNA is processed. The processor uses Field Programable Gate Arrays (FPGA) which can be reconfigured depending on the specific application.
He explained that while an Intel processor runs everything in sequence, thus sometimes wasting computer time, the FPGA runs only those specific sections needed. Where there are smaller data sets, the Intel may be swifter.
He gave the example of a Smith-Waterman parallelization (a form of computation that might be used in DNA calculation) where normally 80% of the time is spent in 20% of the calculation. The FPGA speeds the process and evens out time spent on each part of the operation.
While the processor is available for a computer's expansion slots, he also told me that there is a USB version: particularly useful for notebook computers in the field.
The final presentation was by Dr. Sissades Tongsima of BIOTEC where there is a Mac cluster with 46 nodes running OS X: a total of 96 CPUs. Storage consists of 23TB Xsan, with more used for mirroring.
He explained that a cluster by itself "will do you no good" so to utilise all nodes effectively, special software is written using the MPI libraries.
He gave an informative and articulate presentation in English concentrating on the uses of the system, especially the way that it was being used in genetic epidemiology: comparing DNA to calculate differences. An example of the value of this might be to discover if genetic makeup could predict responses to drugs.
One current project was looking at incidences of thalassemia (a hereditary blood disease caused by faulty hemoglobin synthesis).
The system was also used to examine how certain proteins interact so that the ways one might inhibit another may be simulated. Dr. Sissades pointed out that Riche used such simulations when developing Tamiflu: protection from the avian flu.
All of Dr Sissades' examples, were situations heavy on calculations. He explained that a multiple cluster will cut down calculations from "millions of weeks" to a few months.
While many of us think of Macs as desktops or portable machines, at the top end, when servers, clusters or massive storage is required, OS X and the high level computers available, particularly with the unlimited licensing that Apple allows can make work easier for the scientific community.
See also this article on MacNewsWorld

For further information, e-mail to
Back to
eXtensions
To
eXtensions: 2004-05
To
eXtensions: Year Two
To
eXtensions: Year One
To
eXtensions: Book Reviews
Back to homepage